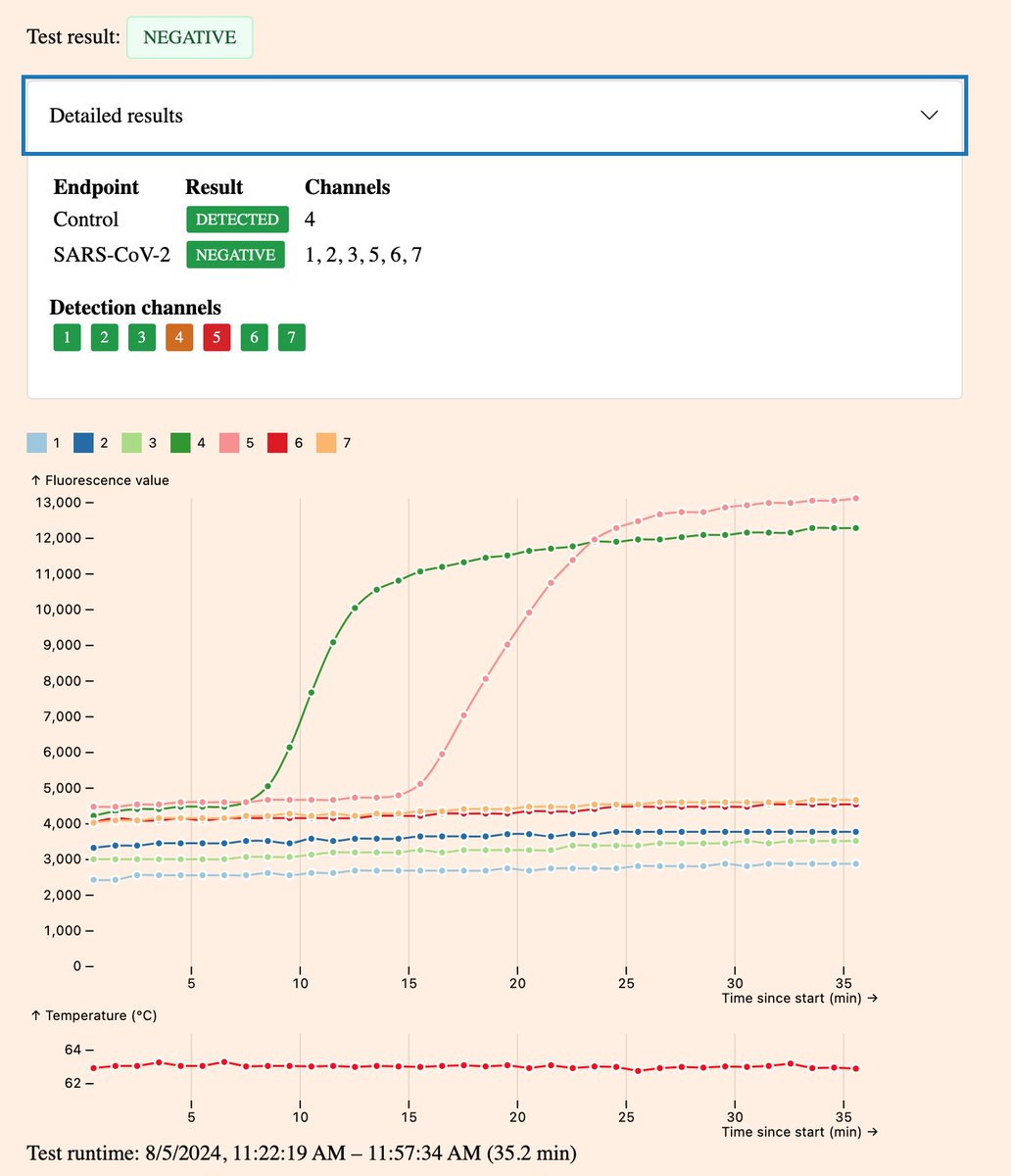
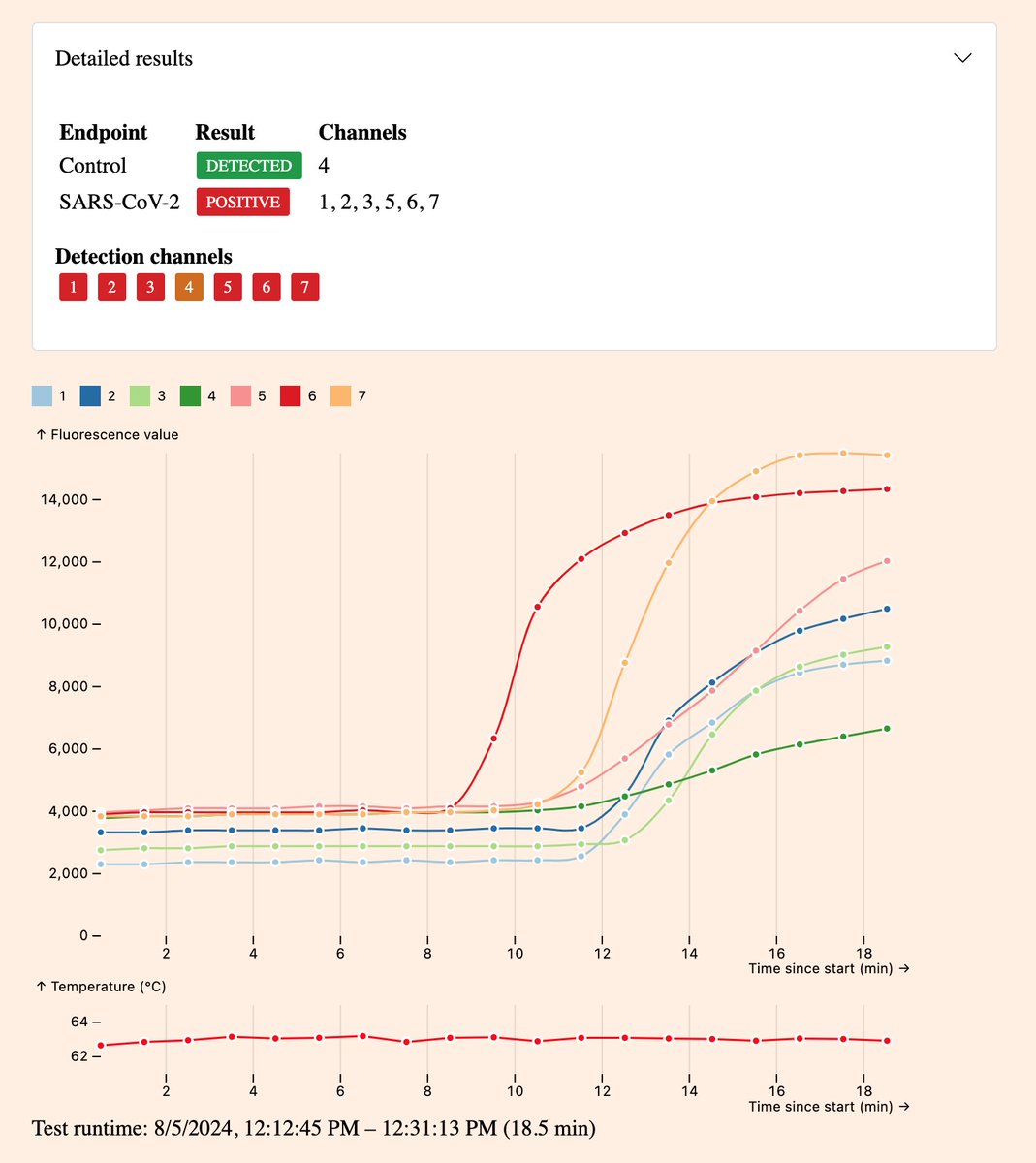
Here’s two different pluslife SARS-CoV-2 results from the same weakly positive sample.
In the first case it was classified as negative as the target was detected in only a single channel. In the second case it was strongly positive as the target was detected in all channels. 1/3
The only difference between these tests is that in the first case I spiked 20µL of my positive sample into a real negative sample from a fresh swab. In the second case I put 20µL of positive sample into buffer on it’s own. 2/3
This highlights a likely problem with lots of PCR sensitivity analysis in the literature: diluting a sample with buffer alone is not a good approximation of a weak sample. Likely the ratio of target to non-target nucleic acid in the sample matters to assay performance! 3/3